Bayesian Neural Networks¶
Neural networks are powerful approximators. However, standard approaches for learning this approximators does not take into account the inherent uncertainty we may have when fitting a model.
import logging, os
logging.disable(logging.WARNING)
os.environ["TF_CPP_MIN_LOG_LEVEL"] = "3"
import matplotlib.pyplot as plt
import numpy as np
import tensorflow as tf
import math
import inferpy as inf
import tensorflow_probability as tfp
Data¶
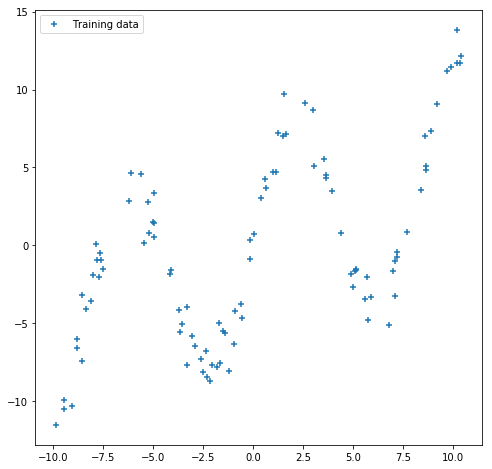
We use some fake data. As neural nets of even one hidden layer can be universal function approximators, we can see if we can train a simple neural network to fit a noisy sinusoidal data, like this:
NSAMPLE = 100
x_train = np.float32(np.random.uniform(-10.5, 10.5, (1, NSAMPLE))).T
r_train = np.float32(np.random.normal(size=(NSAMPLE,1),scale=1.0))
y_train = np.float32(np.sin(0.75*x_train)*7.0+x_train*0.5+r_train*1.0)
plt.figure(figsize=(8, 8))
plt.scatter(x_train, y_train, marker='+', label='Training data')
plt.legend();
Learning Standard Neural Networks¶
We employ a simple feedforward network with 20 hidden units to learn the model.
NHIDDEN = 20
nnetwork = tf.keras.Sequential([
tf.keras.layers.Dense(NHIDDEN, activation=tf.nn.tanh),
tf.keras.layers.Dense(1)
])
lossfunc = lambda y_out, y: tf.nn.l2_loss(y_out-y)
nnetwork.compile(tf.train.AdamOptimizer(0.01), lossfunc)
nnetwork.fit(x=x_train, y=y_train, epochs=1000)
Epoch 1/1000
100/100 [==============================] - 0s 2ms/sample - loss: 369.0472
Epoch 2/1000
100/100 [==============================] - 0s 110us/sample - loss: 333.4858
Epoch 3/1000
100/100 [==============================] - 0s 58us/sample - loss: 339.3102
[...]
Epoch 997/1000
100/100 [==============================] - 0s 52us/sample - loss: 78.6340
Epoch 998/1000
100/100 [==============================] - 0s 50us/sample - loss: 78.3568
Epoch 999/1000
100/100 [==============================] - 0s 37us/sample - loss: 78.4461
Epoch 1000/1000
100/100 [==============================] - 0s 49us/sample - loss: 76.3849
<tensorflow.python.keras.callbacks.History at 0x139726668>
We see that the neural network can fit this sinusoidal data quite well, as expected.
sess = tf.keras.backend.get_session()
x_test = np.float32(np.arange(-10.5,10.5,0.1))
x_test = x_test.reshape(x_test.size,1)
y_test = sess.run(nnetwork(x_test))
plt.figure(figsize=(8, 8))
plt.plot(x_test, y_test, 'r-', label='Predictive mean');
plt.scatter(x_train, y_train, marker='+', label='Training data')
plt.xticks(np.arange(-10., 10.5, 4))
plt.title('Standard Neural Network')
plt.legend();

However, the model uncertainty is not appropriately captured. For example, when making predictions about a single point (e.g. around x=2.0), we can see we do not have into account the inherent noise there is in the prediction. In the next section, we will what happen when we introduce a Bayesian approach using InferPy.
Learning Bayesian Neural Networks¶
Bayesian modeling offers a systematic framework for reasoning about model uncertainty. Instead of just learning point estimates, we’re going to learn a distribution over variables that are consistent with the observed data.
In Bayesian learning, the weights of the network are
random variables. The output of the nework is another
random variable which is the one that implicitlyl defines the
loss function. So, when making Bayesian learning we do not define
loss functions, we do define random variables. For more
information you can check this
talk
and this paper.
In Inferpy, defining a Bayesian
neural network is quite straightforward. First we define the neural
network using inf.layers.Sequential and layers of class
tfp.layers.DenseFlipout. Second, the input x and output y
are also defined as random variables. More precisely, the output y
is defined as a Gaussian random varible. The mean of the Gaussian is the
output of the neural network.
@inf.probmodel
def model(NHIDDEN):
with inf.datamodel():
x = inf.Normal(loc = tf.zeros([1]), scale = 1.0, name="x")
nnetwork = inf.layers.Sequential([
tfp.layers.DenseFlipout(NHIDDEN, activation=tf.nn.tanh),
tfp.layers.DenseFlipout(1)
])
y = inf.Normal(loc = nnetwork(x) , scale= 1., name="y")
To perform Bayesian learning, we resort to the scalable variational
methods available in InferPy, which require the definition of a q
model. For details, see the documentation about Inference in
Inferpy.
For a deeper theoretical despcription, read this
paper. In this case, the q
variables approximating the NN are defined in a transparent manner. For
that reason we define an empty q model.
@inf.probmodel
def qmodel():
pass
NHIDDEN=20
p = model(NHIDDEN)
q = qmodel()
VI = inf.inference.VI(q, optimizer = tf.train.AdamOptimizer(0.01), epochs=5000)
p.fit({"x": x_train, "y": y_train}, VI)
0 epochs 3477.63818359375....................
200 epochs 2621.487548828125....................
400 epochs 2294.40478515625....................
600 epochs 2003.2978515625....................
800 epochs 1932.5308837890625....................
1000 epochs 1912.515625....................
1200 epochs 1909.4072265625....................
1400 epochs 1908.7269287109375....................
1600 epochs 1908.28564453125....................
1800 epochs 1909.939697265625....................
2000 epochs 1907.779052734375....................
2200 epochs 1908.8096923828125....................
2400 epochs 1907.308349609375....................
2600 epochs 1907.8809814453125....................
2800 epochs 1906.529541015625....................
3000 epochs 1906.2943115234375....................
3200 epochs 1906.744140625....................
3400 epochs 1905.798828125....................
3600 epochs 1905.2296142578125....................
3800 epochs 1905.57275390625....................
4000 epochs 1905.6163330078125....................
4200 epochs 1904.5223388671875....................
4400 epochs 1904.778564453125....................
4600 epochs 1904.68408203125....................
4800 epochs 1903.94970703125....................
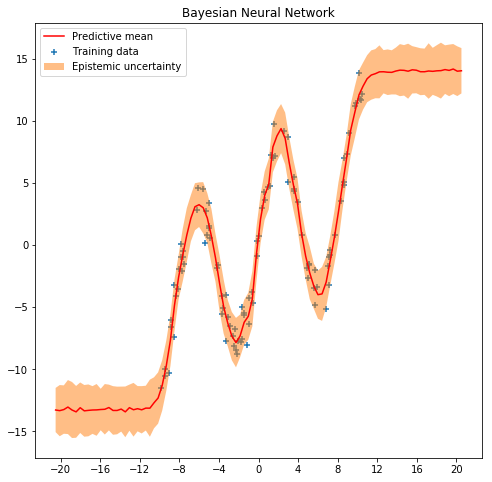
As can be seen in the next figure, the output of our model is not
deterministic. So, we can capture the uncertainty in the data. See for
example what happens now with the predictions at the point x=2.0.
See also what happens with the uncertainty in out-of-range predictions.
x_test = np.linspace(-20.5, 20.5, NSAMPLE).reshape(-1, 1)
plt.figure(figsize=(8, 8))
y_pred_list = []
for i in range(100):
y_test = p.posterior_predictive(["y"], data = {"x": x_test}).sample()
y_pred_list.append(y_test)
y_preds = np.concatenate(y_pred_list, axis=1)
y_mean = np.mean(y_preds, axis=1)
y_sigma = np.std(y_preds, axis=1)
plt.plot(x_test, y_mean, 'r-', label='Predictive mean');
plt.scatter(x_train, y_train, marker='+', label='Training data')
plt.fill_between(x_test.ravel(),
y_mean + 2 * y_sigma,
y_mean - 2 * y_sigma,
alpha=0.5, label='Epistemic uncertainty')
plt.xticks(np.arange(-20., 20.5, 4))
plt.title('Bayesian Neural Network')
plt.legend();